Introduction
Take a moment to consider that there are ten times more bacteria present in the human gut than there are human cells in the body. Surprising and shocking as this may be, it should also occur to you that such vast numbers of bacteria are not there just by chance. In fact, these populations play numerous vital roles in our health and daily functioning. There are at least 100 trillion bacterial cells in the human gut, comprising over 500 different types, and these bacteria are involved in diverse and vital roles such as the digestion of foods, including foods which we would otherwise be unable to metabolise due to a lack of appropriate enzymes. These gut bacteria also contribute to the development of the gut-associated lymphoid tissue (GALT; part of the immune system located in the gut which is vital for developing tolerance to beneficial bacteria). Additionally, these gut bacteria synthesize vitamins (B and K) and aid in the elimination of harmful toxins.
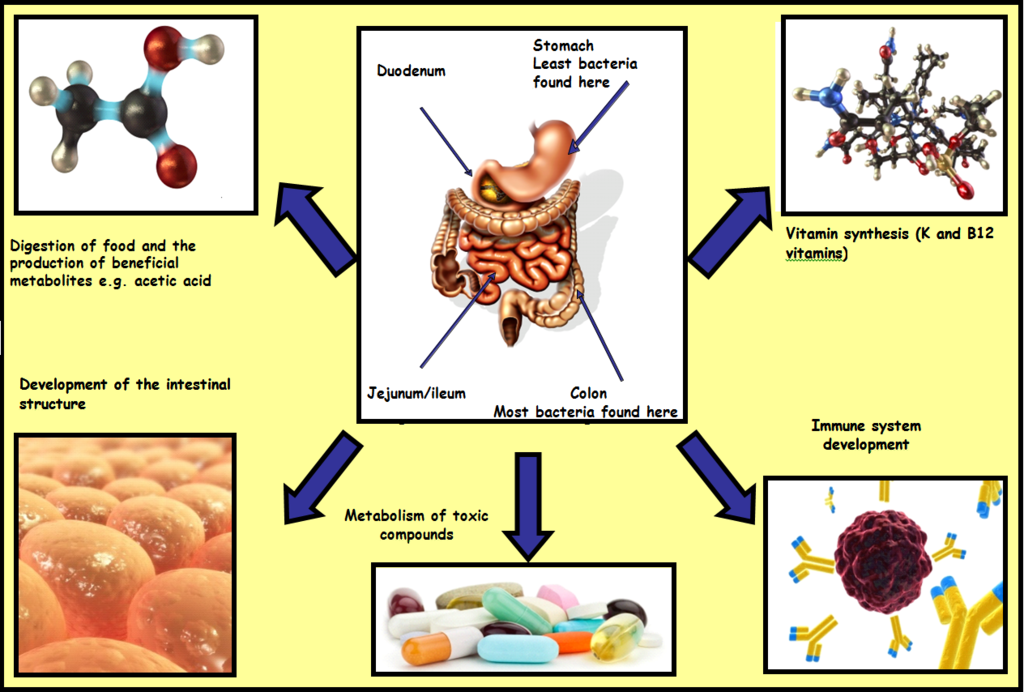
So where do all of these bacteria come from? The process of acquiring these bacteria begins immediately at birth. During delivery, the infant gut is rapidly colonised with bacteria from their mother (if birth is by vaginal delivery) or from hospital staff and the surrounding environment (when birth is by Caesarean delivery). From birth through to 2 years of age, the infant gut undergoes dramatic acquisition, development and shifts in gut bacterial populations. By 2 years of age, the infant’s gut bacterial composition resembles that of an adult, and as such becomes less dynamic, more complex and more stable.
During infancy, many factors other than mode of delivery also affect the bacteria that the infant acquires and develops. These include:
- Feeding choice (breast vs. formula feeding)
- Number of siblings and home environment (e.g. farm vs. urban setting)
- Time of weaning and choice of foods
- Exposure to probiotics (i.e. bacteria which when consumed in foods as part of a healthy diet could help to beneficially colonise the gut conveying measurable health effects)
- Exposure to prebiotics (types of carbohydrates added to foods which act as a nutrient source for beneficial bacteria in the gut, thus increasing their populations)
However, the infant gut bacteria populations can also be negatively affected by exposure to antibiotics in early life. While antibiotics have been pivotal in the prolongation of life, the use of broad spectrum antibiotics (those which kill a wide range of different bacteria) can also cause collateral damage and thus can kill beneficial bacteria such as bifidobacteria and lactobacilli (i.e. those most commonly studied for their health promoting properties). This could potentially increase the risk of the infant developing both short-term and long-term health effects, including antibiotic-associated diarrhoea or, more worryingly, having an increased risk of developing atopic diseases (e.g. eczema or asthma). Antibiotic-treated infants may also be at an increased risk of becoming overweight or obese, an issue causing concern in the current climate. Therefore understanding the exact effects of different antibiotics on both the potentially harmful, as well as the beneficial bacteria in the gut is vital to allow us to fight infections effectively while also minimising the harmful effects on beneficial bacteria.
Study Aims
To date research on the affects of early life exposure to antibiotics on the infant gut bacteria populations has been extremely limited, with much of the research having used culture (or growth)-based approaches. Furthermore, to date, no study has used the new and sophisticated sequencing techniques available to investigate the effects of antibiotics on the gut microbiota of infants.
Thus, the aim of our study was to investigate the effects of treatment with a combination of 2 broad spectrum antibiotics (ampicillin and gentamycin) within 48 hours of birth (for a variety of different reasons) on the gut microbiota of 9 infants compared to 9 healthy controls. Controls are infants who are similar, in all respects, to the antibiotic-treated infants, except they were not exposed to antibiotics in early life. They therefore represent the gut bacteria populations you expect to see in a healthy infant who has not received antibiotics.
Approach Taken
One challenge that exists when studying the gut bacteria populations is accessing a representative sample to study. Most often, a faecal sample is collected. However, this is not ideal, in that, although it is an accurate representative of the bacteria present in the colon (the portion of the gut most densely populated with bacteria), it does not reflect the composition of the bacteria in the upper gastrointestinal tract. Nevertheless, faecal samples are a useful means of studying the gut bacteria populations, are provided in a non-invasive manner and thus continue to be frequently used in the majority of studies, including this present study.
In the past the most common approach was to dilute the faecal samples and grow the bacteria on agar, a jelly like material, and study the resulting growth. However, such approaches are extremely limited as 70-90% of bacteria in the gut do not grow well (or at all) on agar or other laboratory growth media. Moving away from growth (or culture)-based approaches, more recently, scientists began to employ molecular DNA-based approaches (so-called culture-independent approaches) to study all of the bacteria present in specific environments (such as the gut) and not just the populations that can be grown in the laboratory. For this approach, total bacterial DNA is extracted from an environment (e.g. faeces) and studied. In the past decade advances in DNA technology (and the development of next generation DNA sequencing technologies) and computing power have resulted in culture-independent approaches becoming even more powerful. Furthermore, these sequencing technologies allow you to not only identify the bacteria present, but also allow you to determine the relative proportions in which they are present also. Such technologies have significantly improved our understanding of complex microbial environments, including the human gut, and thus the interest in this field of science has increased greatly. However, to date these approaches have been relatively under-utilized in studying the bacteria present in the infant gut. Therefore, my study aimed to use these new technologies to provide detailed insights into the affects of antibiotics in early life on the bacteria present in the gut of infants.
For my study, faecal samples were taken 4 and 8 weeks after antibiotic treatment had ceased. DNA was extracted from these samples and prepared for sequencing. Results were then analysed using bioinformatics, which combines sophisticated computer science with statistical analysis. This approach allowed us to gain a comprehensive insight into the effects of antibiotics on bacterial populations in the infant gut.
Results
When the results from the treated infants were compared to those of the controls, 4 weeks after treatment had ceased, the results were striking.
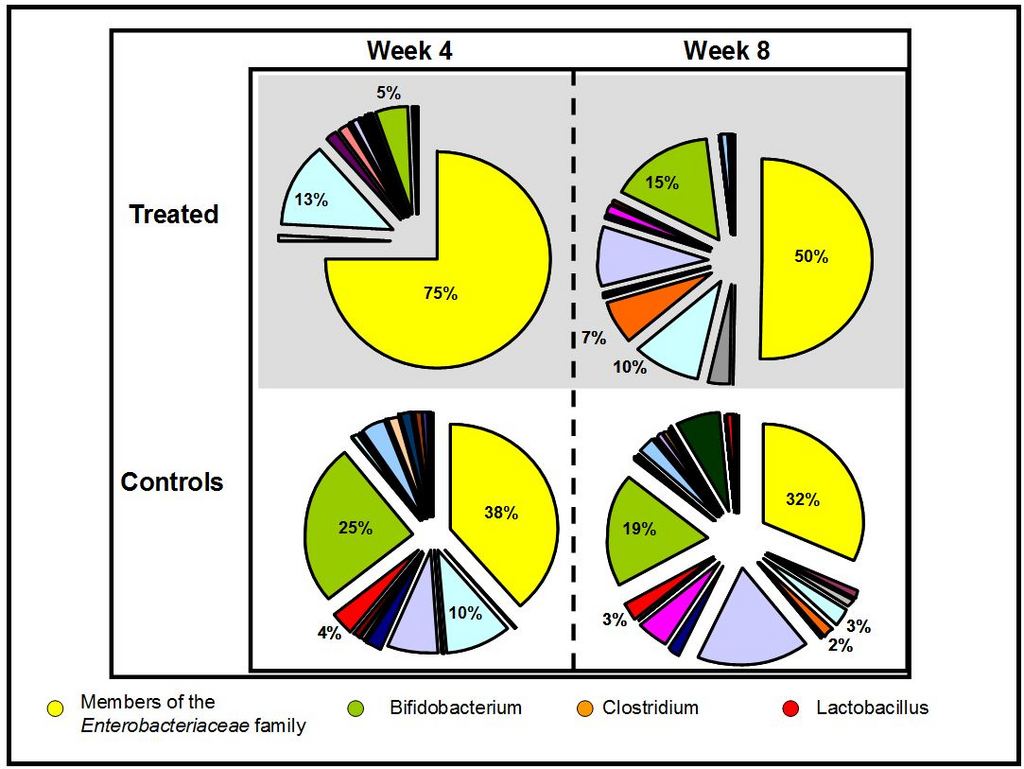
The results indicated that the predominant bacteria present in the antibiotic-treated infants at week 4 were members of the Enterobacteriaceae family e.g. E. coli and Klebsiella (Figure 1). These bacteria dominated the gut of the treated infants, accounting for 75% of all bacteria detected. In contrast, the healthy controls had much lower levels of these potentially harmful bacteria (E. coli and Klebsiella) (38% combined), as well as much higher levels of beneficial bifidobacteria (25%) compared to the treated samples (5%). Lactobacillus levels were also higher in the controls (4%) compared to the treated infants (<1%). The total number of bacteria species detected in the controls was higher than those of the antibiotic-treated samples, who had much less diverse gut bacteria populations, as the majority of gut bacteria were reduced or removed following treatment.
Interestingly, 8 weeks after antibiotic treatment had ceased, there were still significant differences in the gut microbial populations of the treated infants compared to the controls, despite some signs of a recovery. Klebsiella and E. coli, as well as other members of the Enterobacteriaceae family remained dominant, accounting for half of all bacteria detected in the treated samples. These bacteria have the ability to cause ill-health if they increase to significantly high enough levels and out number the beneficial bacteria in the gut. Therefore, even 8 weeks after antibiotic treatment finished, the infant may still be at risk of adverse side effects because of these changes to the gut bacteria.
Levels of beneficial bifidobacteria had recovered so that, 2 months after treatment with antibiotics had ceased, the levels seen in the treated infants were no longer significantly different compared to the controls. This means that despite the infant’s gut being immature, the beneficial bacteria present still have the ability to recover after antibiotic treatment, albeit a somewhat slow recovery of up to 8 weeks. In the controls at week 8, a much more diverse and varied gut microbial population was still evident compared to the treated infants, with no one bacteria being dominant. This showed that not all bacteria could recover over the study period of 8 weeks, and thus even short-term (2-9 days) antibiotic exposure had significant effects on the gut bacteria present.
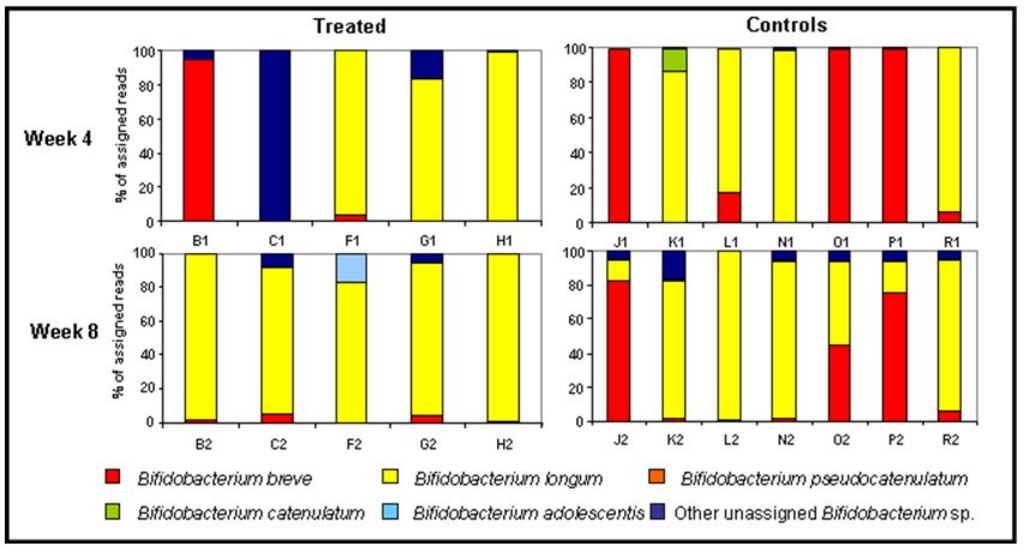
Bifidobacteria have received significant attention as potential beneficial bacteria, which when consumed in foods e.g. yoghurts may have beneficial effects on health. Therefore, it is believed that maintaining good levels of bifidobacteria in the gut is important for health. Given that they have been seen to decrease in the gut following antibiotic treatment, we decided to specifically examine the effects of antibiotic treatment on beneficial bifidobacteria in the infant gut. This study found that only one type of bifidobacteria, B. longum, was dominant following treatment, showing that even within a group of bacteria, not all members will behave in a similar manner following antibiotic treatment (Figure 2).
Conclusions
Although the infant gut is immature until the age of 2 years, it is still a complex and highly dynamic microbial ecosystem. In fact, some have described the gut bacteria populations as a super organ within us, such is the complexity and importance of these bacteria. These gut bacteria are vital to infant health, and changes in levels of beneficial bacteria could have implications on both short-term and long-term health outcomes. Recent research suggests that an altered gut microbiota in early life could predispose the infant to allergic diseases or becoming overweight, obese or developing diabetes in later life.
Our study is the first to demonstrate, using state of the art sequencing technology, the devastating short-term effects of antibiotic therapy on the infant gut microbiota.
The results demonstrate the ability of certain potentially harmful bacteria e.g. E. coli to withstand certain antibiotic treatment and to become the dominant bacteria present. Moreover, the results showed the detrimental effects of antibiotics on levels of beneficial bifidobacteria and lactobacilli and that recovery was slow, with effects still being measureable 8 weeks after treatment had ceased.
Knowing the role of these gut bacteria in the immune system’s development, such dramatic shifts in composition in early life could alter the immune development and potentially predispose the antibiotic treated infant to allergic diseases or even obesity in later life. With such diseases on the increase, understanding the multiple factors involved is essential to help us tackle these diseases effectively. Given the results of our study and that of others previously, it appears that antibiotics are a considerable factor in the development of the gut bacteria populations in infants and that the disturbances they cause could have the potential to cause serious health effects.
So what can be done? Do we continue to prescribe antibiotics to infants and let the beneficial bugs fight for their lives? Based on my findings it may be beneficial to prescribe the use of probiotics (beneficial bacteria in a tablet form or in foods) during antibiotic therapy and also continued use for a period after antibiotic therapy ceases. These bacteria may help to repopulate the infant’s gut with beneficial bacteria populations and thus limit short-term health effects such as gastrointestinal (digestive system) upset and antibiotic-associated diarrhoea and also the long-term effects previously outlined. Additionally, where possible the use of narrow spectrum antibiotics should be used to minimise the collateral damage associated with broad spectrum antibiotic usage, as highlighted in this present study. Thus, while antibiotic use is vital, a targeted approach may be most beneficial to the gut bacterial populations.
The results of this study are important, as using new sophisticated technologies we have shown the significant effects that antibiotics can have on the gut bacteria present in the infant gut. Our approach of sequencing techniques, allowed us to get much more detailed results than if we had used the older growth-based approaches. The results highlight the need to use antibiotics which have a narrow spectrum of activity (i.e. they kill specific bacteria) where possible and the potential benefits of using probiotics to replenish beneficial gut bacteria populations.
This study adds substantially to our understanding of the effects that external factors such as antibiotics can have on the complex gut microbiota environment. Undoubtedly, the insights that are possible through the application of next generation sequencing to studying the gut microbiota, will fuel both scientists’ and the publics’ interest in the marvellous microbial world within us.
Acknowledgments: Fiona Fouhy would like to acknowledge funding received as an EMBARK scholarship from the Irish Research Council for Science, Engineering and Technology and also a Teagasc Walsh fellowship as well as acknowledging the advice and support provided by her supervisors Dr. Paul Cotter, Dr. Catherine Stanton, Prof. Paul Ross and Prof. Gerald Fitzgerald. Additionally the author wishes to acknowledge the contribution of the following individuals to the research contained in this article: Dr. Caitríona Guinane, Dr. Séamus Hussey, Dr. Rebecca Wall, Prof. Anthony Ryan, Dr. Gene Dempsey and Dr. Brendan Murphy.