Bacteriocins
In order for bacteria to survive and grow in a particular environment, they must be able to defend themselves against competing bacteria. To do this, many produce antimicrobial compounds which kill their competitors without harming themselves. Bacteriocins are an excellent example of bacterially produced antimicrobials. Bacteriocins produced by lactic acid bacteria, which are safe bacteria found in milk, cheese and a variety of fermented foods, are the focal point of this research.
Nisin is by far the most researched of all bacteriocins. Its first use as a food preservative was in 1953 and its use has been approved in 48 counties worldwide. After nisin, lacticin 3147 is one of the most thoroughly researched bacteriocins. The bacterial strain that produced lacticin 3147 was found in Ireland and lacticin 3147 is an Irish-owned bacteriocin. Although there are today a number of other bacteriocins available, nisin and lacticin 3147 are the primary interests of this research.
Extensive research into bacteriocins has confirmed them to be safe and natural food antimicrobials that can kill a number of disease-causing, as well as spoilage bacteria, in natural yoghurt, cheese and a variety of other foods. In addition, an exciting application of bacteriocins lies in their potential as antimicrobials against pathogens (disease-causing bacteria) responsible for hospital-associated infections such as methicillin-resistant Staphylococcus aureus (MRSA) and Clostridium difficile (Cdiff).
Mycobacterium avium subspecies paratuberculosis
Although Mycobacterium avium subspecies paratuberculosis (MAP) is not as commonly known as some other disease-causing bacteria, it has gained much attention in recent years due to its association with Crohn’s disease. Along with ulcerative colitis, this disease is categorised as an Inflammatory Bowel Disorder (IBD). Crohn’s disease is a chronic inflammation of the gastrointestinal tract and patients with this disease often present with severe abdominal pains, rectal bleeding, arthritis, diarrhoea and ulceration of the stomach. IBD, in general, effects males and females equally, with those aged between 15 to 25 being most at risk. The question of whether a single type of bacteria could be responsible for this condition has yet to be answered. Conflicting scientific publications have lead to ambiguous viewpoints which continually keep the question of association open.
There is no doubt, however, regarding the bacteria’s role in Johne’s disease. Johne’s disease is a contagious disease which can affect all ruminant animals. There are four unique stages of infection: silent infection, subclinical, clinical and advanced clinical disease. The disease manifests itself through localized chronic infections of the lower intestine (ileum) which cause a thickening of the cell wall. This prevents the normal uptake and absorption of nutrients, which in turn leads to a characteristic wasting away of the animal, despite the fact that the animal seems to be eating very healthily. As the infection increases, the lack of protein uptake causes a condition know as “bottle jaw” where an obvious swelling occurs under the jaw. Infected cattle become increasingly emaciated and die of dehydration and severe cachexia (defined as fatigue and weight loss in an individual/animal that is not actively trying to lose weight).
Unfortunately, MAP can be excreted into the milk and faeces of infected animals in very high numbers which opens a possible route for human infection. Concern regarding the survival of the bacteria in pasteurised milk has also raised more questions than it has answered as conflicting scientific publications do not agree on the effectiveness of this process ineliminating MAP. Although the levels of MAP found in milk and milk products at retail level is extremely low, the bacterium is very difficult to grow in the laboratory and thus numbers present can be dramatically underestimated. Due to the similarity between the symptoms of Johne's and Crohn's diseases, and until it can be definitively established that the consumption of MAP infected milk does not lead to human infections, the elimination of this pathogen from the food chain is highly desirable.
Difficulties with respect to growing MAP is just one of a number of challenges which face MAP researchers. The rate at which MAP grows is exquisitely slow in comparison to other bacteria. For example, a population of a typical laboratory bacteria (E. coli) grows in about 16 hours whereas MAP can take an incredible 16 weeks to grow to detectable levels. Furthermore, the external layer of the bacteria cell is extremely thick, which makes it resistant to a number of forms of antibiotics and disinfectants including chlorine. Even though these characteristics can be seen as an obstacle, they also offer a unique challenge for researchers who wish to gain a greater insight into the world of Mycobacterium avium subspecies paratuberculosis.
Aims of this research
The aim of this research is to take advantage of the antimicrobial activity of bacteriocins by identifying bacteriocins that have particularly good anti-Mycobacterium activity (can kill or inhibit the proliferation of Mycobacteria) and using these in food.
As noted earlier, the rate at which MAP grows is extremely slow. Therefore another Mycobacterium, the non-pathogen Mycobacterium smegmatis, is often used for preliminary investigations. The first goal of this project was to screen a collection of tens of thousands of bacteriocin producers (each producing slightly different forms of nisin) to identify those which produce forms of the bacteriocins which most efficiently kill Mycobacterium smegmatis, with a view to then testing the activity of these compounds against MAP. The bacteriocin-producing bacteria that produce the best results will then be selected and the bacteriocin being produced will become the subject of closer investigation. This requires the purification of the bacteriocin, after which the ability of the purified bacteriocin powder to kill Mycobacteium smegmatis and MAP will be assessed more specifically.
Unfortunately the bacteria within our bacteriocin-producing collection are genetically modified (GM) microorganisms and thus are not permitted in food even if they efficiently kill MAP. However we have at our disposal a technology which allows us to generate non-GMM forms of the most interesting bacteriocin-producing bacteria. Thus bacteria producing these lead-peptides will be recreated using a food-grade approach, followed by the production of a food-grade powder that can be assessed to test its ability to control MAP in food.
Results so far
Since this project commenced in Feb 2009, one of the first successes of this project has been the implementation of rapid screening approach to quickly assess the activity of bacteriocin-producing bacteria against the rapidly growing mycobacterium, Mycobacterium smegmatis. This was achieved by growing the bacteriocin-producing bacteria as localised ‘spots’ on agar plates which hopefully will produce the bacteriocin in the region encircling the ‘spot’. The Mycobacterium is then added but if the bacteriocin has anti-Mycobacterium activity (i.e., it produces a chemical/protein which has the potential to kill the bacteria), the Mycobacterium will not be able to grow around the ‘spot’ and a zone of clearing will be seen (see Figure 1). If the zone of clearing is greater than that produced by nisin or lacticin 3147, then it is apparent that we have identified an improved alternative to these bacteriocins which can be the focus of further investigation. Once this procedure was optimized, rapid screening revealed that a number of strains producing forms of Nisin that inhibit M. smegmatis more effectively than the existing form of the bacteriocin (see Fig. 1). Analysis of the improved forms of nisin revealed that they had enhanced antimicrobial activity as a consequence of genetic changes corresponding with a particular region of the structure of the nisin molecule known as the ‘hinge’. A number of these ‘hinge’-altered nisin proteins will be the focus of further study.
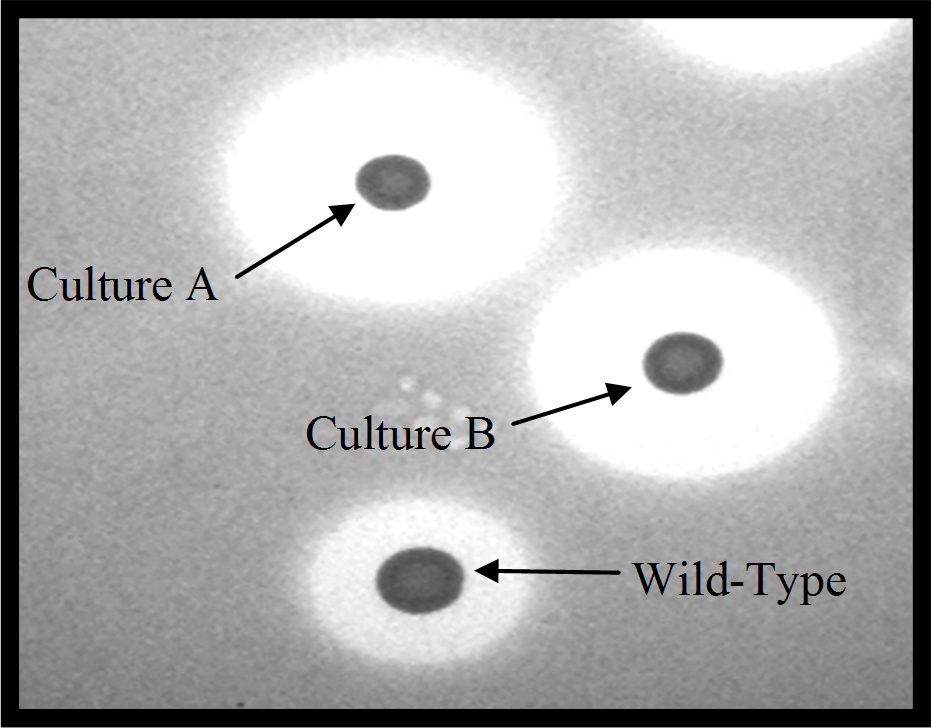
From the picture it can be clearly seen that the zone of clearing for cultures A and B, which are both new forms of nisin,
are far greater than that produced by the original nisin producer [wild-type]
In addition to the existing bank of bacteria producing nisin-like bacteriocins, a similarly sized bank of bacteria producing lacticin 3147-like bacteriocins is in the process of being created and again this bank will be screened, with the lead bacteriocins going forward for further assessment.
In parallel with these studies, another issue being addressed is the question of whether the bacterium can survive the cheese-making process or more specifically the manufacture of a smear ripened cheese produced from raw milk. The smear is a collection of various bacteria, yeasts and moulds deliberately added to the surface of the cheese which gives the cheese its specific taste and texture. This on-going study is also investigating the ability of a lacticin 3147-producing cheese starter culture to limit the survival of MAP. To facilitate an accurate assessment of the survival of MAP both agar-based and molecular methods (methods which specifically target the genetic make-up of the organism), known as Real-Time Polymerase Chain Reaction, will be employed to detect MAP. Hurdles to be overcome in this study relate to the levels of background bacteria, yeasts and moulds other than MAP, in raw milk, the cheese smear, which interferes with the growth characteristics of MAP and the optimisation of methods to extract the bacterial DNA from the dairy products.
Conclusions
The frequency with which new nisin bacteriocins with anti-Mycobacterium activity have been discovered has been a cause of great excitement. With additional collections of producers of nisin-like bacteriocins to be screened, and the potential of lacticin 3147-like bacteriocins yet to be fully realised with respect to MAP, it is anticipated that a large arsenal of anti-MAP compounds for food applications will soon be at our disposal. For more detailed reviews regarding bacteriocins, and their food and clinical applications, see Bacteriocins: developing innate immunity for food, in Nature Reviews Microbiology(Cotter et al, 2005) and Discovery of medically significant lantibiotics in Current Drug Discovery Technologies (Piper et al, 2009).
Brian Healy is a student of the Microbiology Department of UCC under the supervision of Prof. Colin Hill, Dr. Paul Cotter and Prof. Paul Ross. The author would like to acknowledge The Department of Agriculture, Fisheries and Food for the funding received to carry out this research.

